Context-Aware MRI Plane Detection
The Ambiguity Problem: Identifying the anatomical plane (Axial, Coronal, Sagittal) of an MRI scan is usually easy—except at the edges. "Near-skull" edge images lack distinct anatomical features, making them inherently ambiguous for AI models and even human experts.
The Consequence: Standard 2D classifiers fail on these ambiguous slices. This leads to corrupted metadata, confusing downstream tasks and causing domain shift when merging heterogeneous datasets.
Our Solution: We introduce a Context-Aware 2.5D Model. By sampling adjacent slices, our model learns the local anatomical flow, providing just enough context to resolve ambiguity. This approach achieves >99% accuracy and corrects misclassifications that purely 2D models cannot solve.
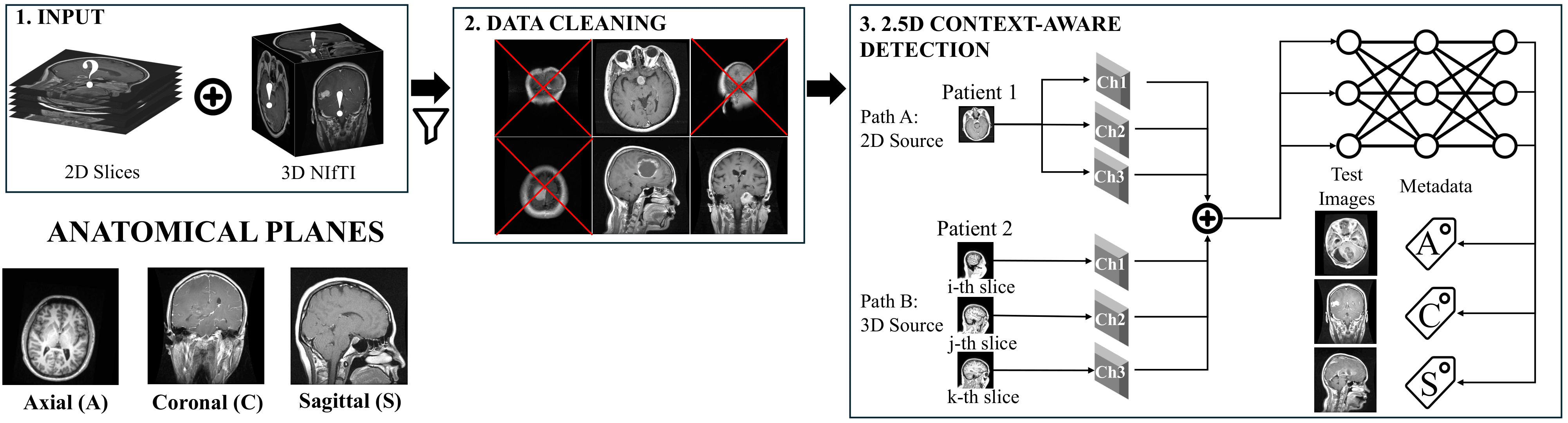
How do we teach a model to "see" context? We construct a 3-channel input where the center slice is the target, flanked by adjacent slices (Sequential) or random slices from the same volume (Random).
We compared a standard 2D baseline against our 2.5D approach on a mixed dataset. The 2.5D model effectively eliminates errors in ambiguous classes.
The chart above shows that the model is better. This visualization shows why. Below shows the "Top-2 Most Confident Errors" made by the 2D model compared to our 2.5D model on the same difficult slices.
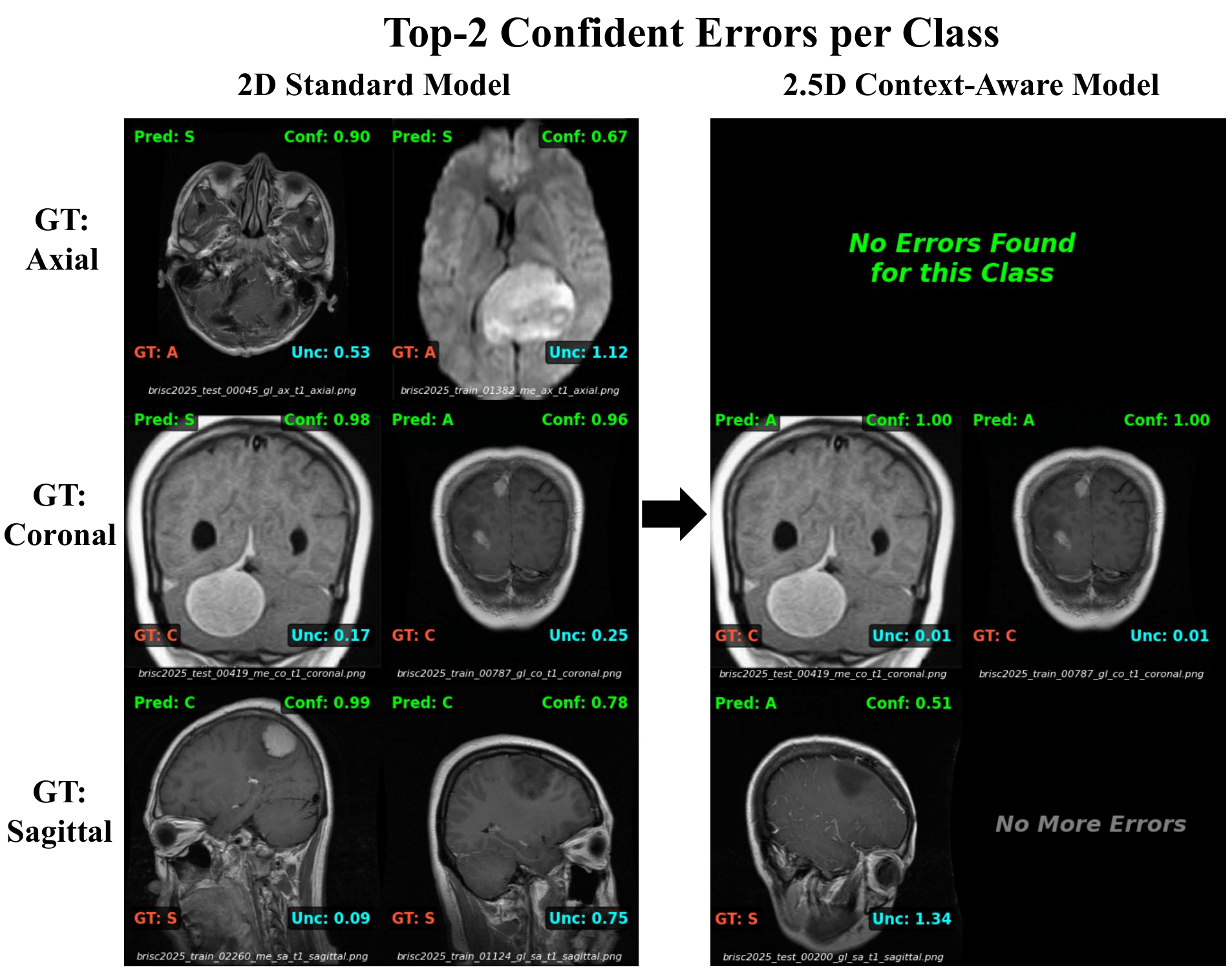
Row 1 (Axial): The 2D model is confused by near-skull slices. The 2.5D model completely eliminates these errors.
Row 2 (Coronal): Large tumors cause asymmetry, fooling the 2D model into predicting "Sagittal". The 2.5D model uses context to correctly identify the plane despite the pathology.
To prove that the model is learning anatomical flow rather than just memorizing images, we trained on 3D volumes (IXI) and tested on an unseen 2D dataset (BRISC).
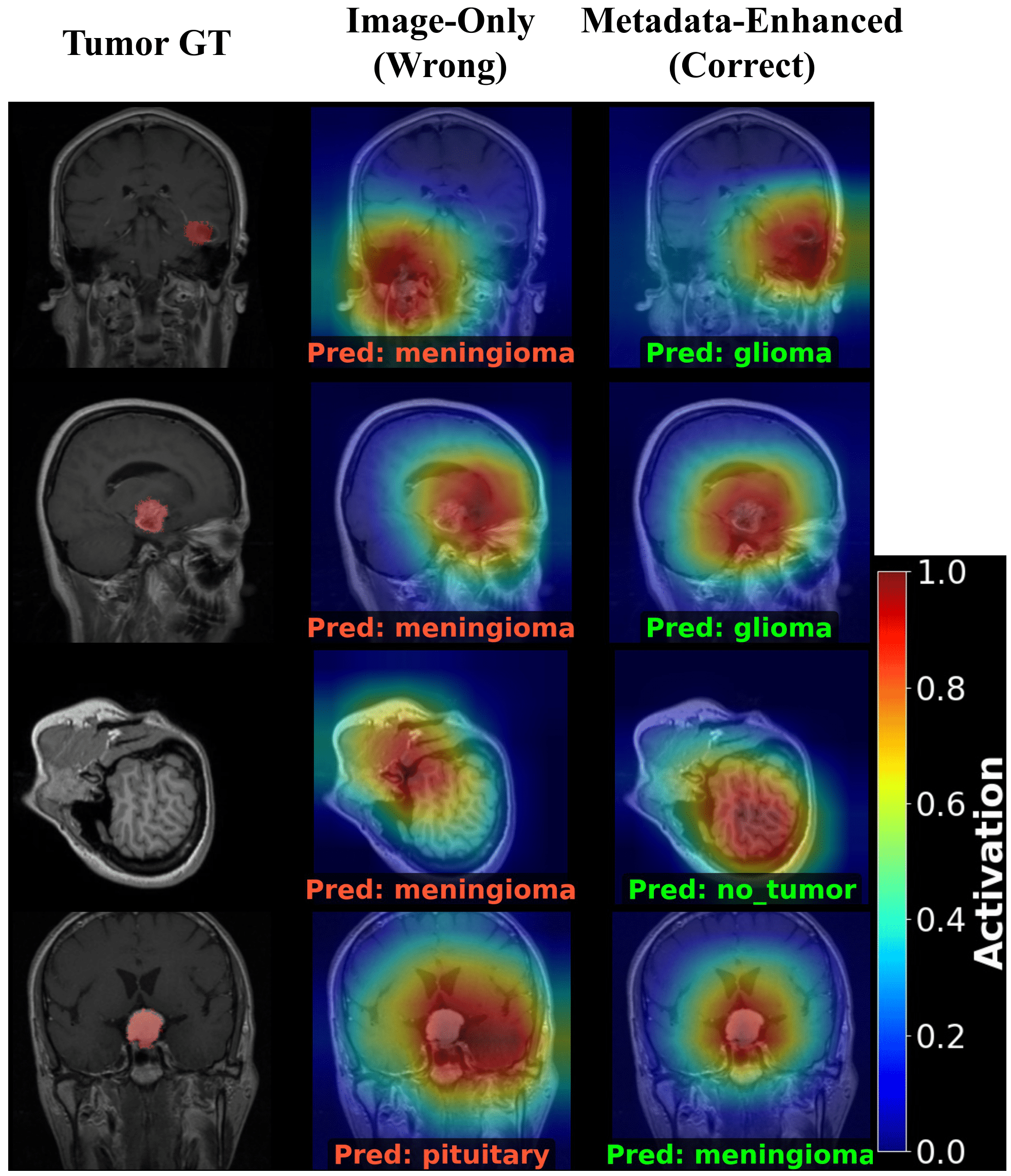
A. Qualitative Analysis (Grad-CAM)
Before looking at the statistics, observe how the corrected metadata fixes the model's behavior. Left: Image-Only (Wrong). Right: Metadata-Enhanced (Correct).
B. Quantitative Impact: Reducing Misdiagnoses
We applied this corrected attention to the full dataset using a Gated Strategy (filtering uncertain predictions).